File formats¶
Miller handles name-indexed data using several formats: some you probably know by name, such as CSV, TSV, and JSON – and other formats you’re likely already seeing and using in your structured data. Additionally, Miller gives you the option of including comments within your data.
Examples¶
$ mlr --usage-data-format-examples
DKVP: delimited key-value pairs (Miller default format)
+---------------------+
| apple=1,bat=2,cog=3 | Record 1: "apple" => "1", "bat" => "2", "cog" => "3"
| dish=7,egg=8,flint | Record 2: "dish" => "7", "egg" => "8", "3" => "flint"
+---------------------+
NIDX: implicitly numerically indexed (Unix-toolkit style)
+---------------------+
| the quick brown | Record 1: "1" => "the", "2" => "quick", "3" => "brown"
| fox jumped | Record 2: "1" => "fox", "2" => "jumped"
+---------------------+
CSV/CSV-lite: comma-separated values with separate header line
+---------------------+
| apple,bat,cog |
| 1,2,3 | Record 1: "apple => "1", "bat" => "2", "cog" => "3"
| 4,5,6 | Record 2: "apple" => "4", "bat" => "5", "cog" => "6"
+---------------------+
Tabular JSON: nested objects are supported, although arrays within them are not:
+---------------------+
| { |
| "apple": 1, | Record 1: "apple" => "1", "bat" => "2", "cog" => "3"
| "bat": 2, |
| "cog": 3 |
| } |
| { |
| "dish": { | Record 2: "dish:egg" => "7", "dish:flint" => "8", "garlic" => ""
| "egg": 7, |
| "flint": 8 |
| }, |
| "garlic": "" |
| } |
+---------------------+
PPRINT: pretty-printed tabular
+---------------------+
| apple bat cog |
| 1 2 3 | Record 1: "apple => "1", "bat" => "2", "cog" => "3"
| 4 5 6 | Record 2: "apple" => "4", "bat" => "5", "cog" => "6"
+---------------------+
XTAB: pretty-printed transposed tabular
+---------------------+
| apple 1 | Record 1: "apple" => "1", "bat" => "2", "cog" => "3"
| bat 2 |
| cog 3 |
| |
| dish 7 | Record 2: "dish" => "7", "egg" => "8"
| egg 8 |
+---------------------+
Markdown tabular (supported for output only):
+-----------------------+
| | apple | bat | cog | |
| | --- | --- | --- | |
| | 1 | 2 | 3 | | Record 1: "apple => "1", "bat" => "2", "cog" => "3"
| | 4 | 5 | 6 | | Record 2: "apple" => "4", "bat" => "5", "cog" => "6"
+-----------------------+
CSV/TSV/ASV/USV/etc.¶
When mlr is invoked with the --csv or --csvlite option, key names are found on the first record and values are taken from subsequent records. This includes the case of CSV-formatted files. See Record-heterogeneity for how Miller handles changes of field names within a single data stream.
Miller has record separator RS and field separator FS, just as awk does. For TSV, use --fs tab; to convert TSV to CSV, use --ifs tab --ofs comma, etc. (See also Record/field/pair separators.)
TSV (tab-separated values): the following are synonymous pairs:
--tsvand--csv --fs tab--itsvand--icsv --ifs tab--otsvand--ocsv --ofs tab--tsvliteand--csvlite --fs tab--itsvliteand--icsvlite --ifs tab--otsvliteand--ocsvlite --ofs tab
ASV (ASCII-separated values): the flags --asv, --iasv, --oasv, --asvlite, --iasvlite, and --oasvlite are analogous except they use ASCII FS and RS 0x1f and 0x1e, respectively.
USV (Unicode-separated values): likewise, the flags --usv, --iusv, --ousv, --usvlite, --iusvlite, and --ousvlite use Unicode FS and RS U+241F (UTF-8 0x0xe2909f) and U+241E (UTF-8 0xe2909e), respectively.
Miller’s --csv flag supports RFC-4180 CSV. This includes CRLF line-terminators by default, regardless of platform.
Here are the differences between CSV and CSV-lite:
CSV supports RFC-4180-style double-quoting, including the ability to have commas and/or LF/CRLF line-endings contained within an input field; CSV-lite does not.
CSV does not allow heterogeneous data; CSV-lite does (see also Record-heterogeneity).
The CSV-lite input-reading code is fractionally more efficient than the CSV input-reader.
Here are things they have in common:
The ability to specify record/field separators other than the default, e.g. CR-LF vs. LF, or tab instead of comma for TSV, and so on.
The
--implicit-csv-headerflag for input and the--headerless-csv-outputflag for output.
DKVP: Key-value pairs¶
Miller’s default file format is DKVP, for delimited key-value pairs. Example:
$ mlr cat data/small
a=pan,b=pan,i=1,x=0.3467901443380824,y=0.7268028627434533
a=eks,b=pan,i=2,x=0.7586799647899636,y=0.5221511083334797
a=wye,b=wye,i=3,x=0.20460330576630303,y=0.33831852551664776
a=eks,b=wye,i=4,x=0.38139939387114097,y=0.13418874328430463
a=wye,b=pan,i=5,x=0.5732889198020006,y=0.8636244699032729
Such data are easy to generate, e.g. in Ruby with
puts "host=#{hostname},seconds=#{t2-t1},message=#{msg}"
puts mymap.collect{|k,v| "#{k}=#{v}"}.join(',')
or print statements in various languages, e.g.
echo "type=3,user=$USER,date=$date\n";
logger.log("type=3,user=$USER,date=$date\n");
Fields lacking an IPS will have positional index (starting at 1) used as the key, as in NIDX format. For example, dish=7,egg=8,flint is parsed as "dish" => "7", "egg" => "8", "3" => "flint" and dish,egg,flint is parsed as "1" => "dish", "2" => "egg", "3" => "flint".
As discussed in Record-heterogeneity, Miller handles changes of field names within the same data stream. But using DKVP format this is particularly natural. One of my favorite use-cases for Miller is in application/server logs, where I log all sorts of lines such as
resource=/path/to/file,loadsec=0.45,ok=true
record_count=100, resource=/path/to/file
resource=/some/other/path,loadsec=0.97,ok=false
etc. and I just log them as needed. Then later, I can use grep, mlr --opprint group-like, etc.
to analyze my logs.
See Main reference regarding how to specify separators other than the default equals-sign and comma.
NIDX: Index-numbered (toolkit style)¶
With --inidx --ifs ' ' --repifs, Miller splits lines on whitespace and assigns integer field names starting with 1. This recapitulates Unix-toolkit behavior.
Example with index-numbered output:
$ cat data/small
a=pan,b=pan,i=1,x=0.3467901443380824,y=0.7268028627434533
a=eks,b=pan,i=2,x=0.7586799647899636,y=0.5221511083334797
a=wye,b=wye,i=3,x=0.20460330576630303,y=0.33831852551664776
a=eks,b=wye,i=4,x=0.38139939387114097,y=0.13418874328430463
a=wye,b=pan,i=5,x=0.5732889198020006,y=0.8636244699032729
$ mlr --onidx --ofs ' ' cat data/small
pan pan 1 0.3467901443380824 0.7268028627434533
eks pan 2 0.7586799647899636 0.5221511083334797
wye wye 3 0.20460330576630303 0.33831852551664776
eks wye 4 0.38139939387114097 0.13418874328430463
wye pan 5 0.5732889198020006 0.8636244699032729
Example with index-numbered input:
$ cat data/mydata.txt
oh say can you see
by the dawn's
early light
$ mlr --inidx --ifs ' ' --odkvp cat data/mydata.txt
1=oh,2=say,3=can,4=you,5=see
1=by,2=the,3=dawn's
1=early,2=light
Example with index-numbered input and output:
$ cat data/mydata.txt
oh say can you see
by the dawn's
early light
$ mlr --nidx --fs ' ' --repifs cut -f 2,3 data/mydata.txt
say can
the dawn's
light
Tabular JSON¶
JSON is a format which supports arbitrarily deep nesting of “objects” (hashmaps) and “arrays” (lists), while Miller is a tool for handling tabular data only. This means Miller cannot (and should not) handle arbitrary JSON. (Check out jq.)
But if you have tabular data represented in JSON then Miller can handle that for you.
Single-level JSON objects¶
An array of single-level objects is, quite simply, a table:
$ mlr --json head -n 2 then cut -f color,shape data/json-example-1.json
{ "color": "yellow", "shape": "triangle" }
{ "color": "red", "shape": "square" }
$ mlr --json --jvstack head -n 2 then cut -f color,u,v data/json-example-1.json
{
"color": "yellow",
"u": 0.6321695890307647,
"v": 0.9887207810889004
}
{
"color": "red",
"u": 0.21966833570651523,
"v": 0.001257332190235938
}
$ mlr --ijson --opprint stats1 -a mean,stddev,count -f u -g shape data/json-example-1.json
shape u_mean u_stddev u_count
triangle 0.583995 0.131184 3
square 0.409355 0.365428 4
circle 0.366013 0.209094 3
Nested JSON objects¶
Additionally, Miller can tabularize nested objects by concatentating keys:
$ mlr --json --jvstack head -n 2 data/json-example-2.json
{
"flag": 1,
"i": 11,
"attributes": {
"color": "yellow",
"shape": "triangle"
},
"values": {
"u": 0.632170,
"v": 0.988721,
"w": 0.436498,
"x": 5.798188
}
}
{
"flag": 1,
"i": 15,
"attributes": {
"color": "red",
"shape": "square"
},
"values": {
"u": 0.219668,
"v": 0.001257,
"w": 0.792778,
"x": 2.944117
}
}
$ mlr --ijson --opprint head -n 4 data/json-example-2.json
flag i attributes:color attributes:shape values:u values:v values:w values:x
1 11 yellow triangle 0.632170 0.988721 0.436498 5.798188
1 15 red square 0.219668 0.001257 0.792778 2.944117
1 16 red circle 0.209017 0.290052 0.138103 5.065034
0 48 red square 0.956274 0.746720 0.775542 7.117831
Note in particular that as far as Miller’s put and filter, as well as other I/O formats, are concerned, these are simply field names with colons in them:
$ mlr --json --jvstack head -n 1 then put '${values:uv} = ${values:u} * ${values:v}' data/json-example-2.json
{
"flag": 1,
"i": 11,
"attributes": {
"color": "yellow",
"shape": "triangle"
},
"values": {
"u": 0.632170,
"v": 0.988721,
"w": 0.436498,
"x": 5.798188,
"uv": 0.625040
}
}
Arrays¶
Arrays aren’t supported in Miller’s put/filter DSL. By default, JSON arrays are read in as integer-keyed maps.
Suppose we have arrays like this in our input data:
$ cat data/json-example-3.json
{
"label": "orange",
"values": [12.2, 13.8, 17.2]
}
{
"label": "purple",
"values": [27.0, 32.4]
}
Then integer indices (starting from 0 and counting up) are used as map keys:
$ mlr --ijson --oxtab cat data/json-example-3.json
label orange
values:0 12.2
values:1 13.8
values:2 17.2
label purple
values:0 27.0
values:1 32.4
When the data are written back out as JSON, field names are re-expanded as above, but what were arrays on input are now maps on output:
$ mlr --json --jvstack cat data/json-example-3.json
{
"label": "orange",
"values": {
"0": 12.2,
"1": 13.8,
"2": 17.2
}
}
{
"label": "purple",
"values": {
"0": 27.0,
"1": 32.4
}
}
This is non-ideal, but it allows Miller (5.x release being latest as of this writing) to handle JSON arrays at all.
You might also use mlr --json-skip-arrays-on-input or mlr --json-fatal-arrays-on-input.
To truly handle JSON, please use a JSON-processing tool such as jq.
Formatting JSON options¶
JSON isn’t a parameterized format, so RS, FS, PS aren’t specifiable. Nonetheless, you can do the following:
Use
--jvstackto pretty-print JSON objects with multi-line (vertically stacked) spacing. By default, each Miller record (JSON object) is one per line.Keystroke-savers:
--jsonxsimply means--json --jvstack, and--ojsonxsimply means--ojson --jvstack.Use
--jlistwrapto print the sequence of JSON objects wrapped in an outermost[and]. By default, these aren’t printed.Use
--jquoteallto double-quote all object values. By default, integers, floating-point numbers, and booleanstrueandfalseare not double-quoted when they appear as JSON-object keys.Use
--jflatsep yourstringhereto specify the string used for key concatenation: this defaults to a single colon.Use
--jofmtto force Miller to apply the global--ofmtto floating-point values. First note: please use sprintf-style codes for double precision, e.g. ending in%lf,%le, or%lg. Miller floats are double-precision so behavior using%f,%d, etc. is undefined. Second note:0.123is valid JSON;.123is not. Thus this feature allows you to emit JSON which may be unparseable by other tools.
Again, please see jq for a truly powerful, JSON-specific tool.
JSON non-streaming¶
The JSON parser Miller uses does not return until all input is parsed: in particular this means that, unlike for other file formats, Miller does not (at present) handle JSON files in tail -f contexts.
PPRINT: Pretty-printed tabular¶
Miller’s pretty-print format is like CSV, but column-aligned. For example, compare
$ mlr --ocsv cat data/small
a,b,i,x,y
pan,pan,1,0.3467901443380824,0.7268028627434533
eks,pan,2,0.7586799647899636,0.5221511083334797
wye,wye,3,0.20460330576630303,0.33831852551664776
eks,wye,4,0.38139939387114097,0.13418874328430463
wye,pan,5,0.5732889198020006,0.8636244699032729
$ mlr --opprint cat data/small
a b i x y
pan pan 1 0.3467901443380824 0.7268028627434533
eks pan 2 0.7586799647899636 0.5221511083334797
wye wye 3 0.20460330576630303 0.33831852551664776
eks wye 4 0.38139939387114097 0.13418874328430463
wye pan 5 0.5732889198020006 0.8636244699032729
Note that while Miller is a line-at-a-time processor and retains input lines in memory only where necessary (e.g. for sort), pretty-print output requires it to accumulate all input lines (so that it can compute maximum column widths) before producing any output. This has two consequences: (a) pretty-print output won’t work on tail -f contexts, where Miller will be waiting for an end-of-file marker which never arrives; (b) pretty-print output for large files is constrained by available machine memory.
See Record-heterogeneity for how Miller handles changes of field names within a single data stream.
For output only (this isn’t supported in the input-scanner as of 5.0.0) you can use --barred with pprint output format:
$ mlr --opprint --barred cat data/small
+-----+-----+---+---------------------+---------------------+
| a | b | i | x | y |
+-----+-----+---+---------------------+---------------------+
| pan | pan | 1 | 0.3467901443380824 | 0.7268028627434533 |
| eks | pan | 2 | 0.7586799647899636 | 0.5221511083334797 |
| wye | wye | 3 | 0.20460330576630303 | 0.33831852551664776 |
| eks | wye | 4 | 0.38139939387114097 | 0.13418874328430463 |
| wye | pan | 5 | 0.5732889198020006 | 0.8636244699032729 |
+-----+-----+---+---------------------+---------------------+
XTAB: Vertical tabular¶
This is perhaps most useful for looking a very wide and/or multi-column data which causes line-wraps on the screen (but see also ngrid for an entirely different, very powerful option). Namely:
$ grep -v '^#' /etc/passwd | head -n 6 | mlr --nidx --fs : --opprint cat
1 2 3 4 5 6 7
nobody * -2 -2 Unprivileged User /var/empty /usr/bin/false
root * 0 0 System Administrator /var/root /bin/sh
daemon * 1 1 System Services /var/root /usr/bin/false
_uucp * 4 4 Unix to Unix Copy Protocol /var/spool/uucp /usr/sbin/uucico
_taskgated * 13 13 Task Gate Daemon /var/empty /usr/bin/false
_networkd * 24 24 Network Services /var/networkd /usr/bin/false
$ grep -v '^#' /etc/passwd | head -n 2 | mlr --nidx --fs : --oxtab cat
1 nobody
2 *
3 -2
4 -2
5 Unprivileged User
6 /var/empty
7 /usr/bin/false
1 root
2 *
3 0
4 0
5 System Administrator
6 /var/root
7 /bin/sh
$ grep -v '^#' /etc/passwd | head -n 2 | \
mlr --nidx --fs : --ojson --jvstack --jlistwrap label name,password,uid,gid,gecos,home_dir,shell
[
{
"name": "nobody",
"password": "*",
"uid": -2,
"gid": -2,
"gecos": "Unprivileged User",
"home_dir": "/var/empty",
"shell": "/usr/bin/false"
}
,{
"name": "root",
"password": "*",
"uid": 0,
"gid": 0,
"gecos": "System Administrator",
"home_dir": "/var/root",
"shell": "/bin/sh"
}
]
Markdown tabular¶
Markdown format looks like this:
$ mlr --omd cat data/small
| a | b | i | x | y |
| --- | --- | --- | --- | --- |
| pan | pan | 1 | 0.3467901443380824 | 0.7268028627434533 |
| eks | pan | 2 | 0.7586799647899636 | 0.5221511083334797 |
| wye | wye | 3 | 0.20460330576630303 | 0.33831852551664776 |
| eks | wye | 4 | 0.38139939387114097 | 0.13418874328430463 |
| wye | pan | 5 | 0.5732889198020006 | 0.8636244699032729 |
which renders like this when dropped into various web tools (e.g. github comments):
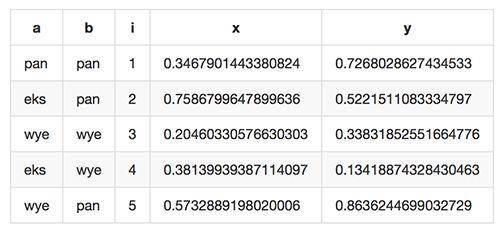
As of Miller 4.3.0, markdown format is supported only for output, not input.
Data-conversion keystroke-savers¶
While you can do format conversion using mlr --icsv --ojson cat myfile.csv, there are also keystroke-savers for this purpose, such as mlr --c2j cat myfile.csv. For a complete list:
$ mlr --usage-format-conversion-keystroke-saver-options
As keystroke-savers for format-conversion you may use the following:
--c2t --c2d --c2n --c2j --c2x --c2p --c2m
--t2c --t2d --t2n --t2j --t2x --t2p --t2m
--d2c --d2t --d2n --d2j --d2x --d2p --d2m
--n2c --n2t --n2d --n2j --n2x --n2p --n2m
--j2c --j2t --j2d --j2n --j2x --j2p --j2m
--x2c --x2t --x2d --x2n --x2j --x2p --x2m
--p2c --p2t --p2d --p2n --p2j --p2x --p2m
The letters c t d n j x p m refer to formats CSV, TSV, DKVP, NIDX, JSON, XTAB,
PPRINT, and markdown, respectively. Note that markdown format is available for
output only.
Autodetect of line endings¶
Default line endings (--irs and --ors) are 'auto' which means autodetect from the input file format, as long as the input file(s) have lines ending in either LF (also known as linefeed, '\n', 0x0a, Unix-style) or CRLF (also known as carriage-return/linefeed pairs, '\r\n', 0x0d 0x0a, Windows style).
If both IRS and ORS are auto (which is the default) then LF input will lead to LF output and CRLF input will lead to CRLF output, regardless of the platform you’re running on.
The line-ending autodetector triggers on the first line ending detected in the input stream. E.g. if you specify a CRLF-terminated file on the command line followed by an LF-terminated file then autodetected line endings will be CRLF.
If you use --ors {something else} with (default or explicitly specified) --irs auto then line endings are autodetected on input and set to what you specify on output.
If you use --irs {something else} with (default or explicitly specified) --ors auto then the output line endings used are LF on Unix/Linux/BSD/MacOSX, and CRLF on Windows.
See also Record/field/pair separators for more information about record/field/pair separators.
Comments in data¶
You can include comments within your data files, and either have them ignored, or passed directly through to the standard output as soon as they are encountered:
$ mlr --usage-comments-in-data
--skip-comments Ignore commented lines (prefixed by "#")
within the input.
--skip-comments-with {string} Ignore commented lines within input, with
specified prefix.
--pass-comments Immediately print commented lines (prefixed by "#")
within the input.
--pass-comments-with {string} Immediately print commented lines within input, with
specified prefix.
Notes:
* Comments are only honored at the start of a line.
* In the absence of any of the above four options, comments are data like
any other text.
* When pass-comments is used, comment lines are written to standard output
immediately upon being read; they are not part of the record stream.
Results may be counterintuitive. A suggestion is to place comments at the
start of data files.
Examples:
$ cat data/budget.csv
# Asana -- here are the budget figures you asked for!
type,quantity
purple,456.78
green,678.12
orange,123.45
$ mlr --skip-comments --icsv --opprint sort -nr quantity data/budget.csv
type quantity
green 678.12
purple 456.78
orange 123.45
$ mlr --pass-comments --icsv --opprint sort -nr quantity data/budget.csv
# Asana -- here are the budget figures you asked for!
type quantity
green 678.12
purple 456.78
orange 123.45